Posterior Predictive Checking with predictCheckR
Utkarsh Pawade
2026-04-04
Source:vignettes/predictCheckR_workflow.Rmd
predictCheckR_workflow.RmdIntroduction
Posterior predictive checking (PPC) is a principled, simulation-based technique for assessing Bayesian model fit. Rather than relying solely on likelihood-based information criteria, PPC asks a direct question:
If the model is correct, would it generate data that look like the data we actually observed?
The Posterior Predictive Distribution
Given a Bayesian model with likelihood and prior , inference yields the posterior . The posterior predictive distribution for a future — or hypothetically replicated — outcome is
In practice the integral is approximated by Monte Carlo averaging over posterior draws :
Each draw is a complete replicated dataset of the same size as . The collection forms the posterior predictive ensemble that is used for visual and numerical diagnostics.
Package Setup
Install predictCheckR from GitHub (once published) or load it from the source directory during development:
The Example Dataset
predictCheckR ships with example_data,
a small simulated regression dataset generated from the model
data(example_data)
head(example_data)
#> x y
#> 1 0.9819694 7.154570
#> 2 0.4687150 3.846278
#> 3 -0.1079713 2.719252
#> 4 -0.2128782 1.487042
#> 5 1.1580985 4.209046
#> 6 1.2923548 6.251425
ggplot(example_data, aes(x = x, y = y)) +
geom_point(alpha = 0.6, colour = "steelblue") +
geom_smooth(method = "lm", se = TRUE, colour = "grey30",
fill = "grey80", linewidth = 0.8) +
labs(
title = "Example Regression Dataset",
subtitle = "True model: y = 2 + 3x + N(0,1)",
x = "Predictor x",
y = "Response y"
) +
theme_ppc()
#> `geom_smooth()` using formula = 'y ~ x'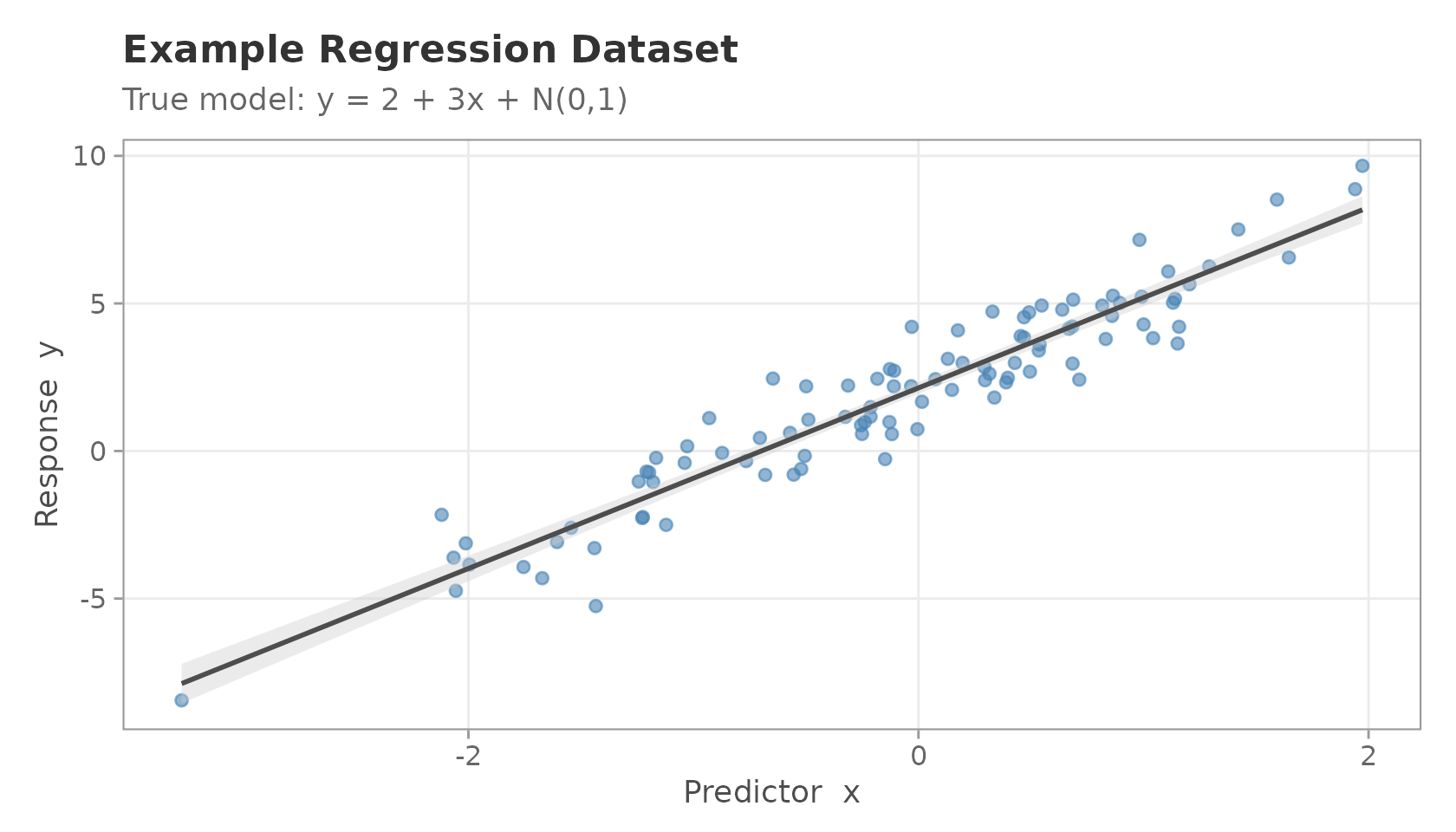
Scatter plot of the example regression dataset.
Generating Fake Posterior Draws
In a real analysis, posterior draws would come from a sampler such as rstan or brms. Here we simulate draws directly to keep the vignette self-contained.
Suppose we have obtained posterior samples of the intercept and slope . We centre the draws on the true values but add realistic posterior uncertainty.
set.seed(101)
S <- 400 # posterior iterations
n <- nrow(example_data)
# Simulate posterior draws: alpha ~ N(2, 0.15^2), beta ~ N(3, 0.12^2)
alpha_draws <- rnorm(S, mean = 2.0, sd = 0.15)
beta_draws <- rnorm(S, mean = 3.0, sd = 0.12)
sigma_draws <- abs(rnorm(S, mean = 1.0, sd = 0.08)) # residual SD draws
# Posterior draws matrix (S x 2): columns are [alpha, beta]
posterior_draws <- cbind(intercept = alpha_draws, slope = beta_draws)
# Design matrix (n x 2): [1, x]
X <- cbind(1, example_data$x)
dim(posterior_draws)
#> [1] 400 2
dim(X)
#> [1] 100 2Generating Posterior Predictive Samples
simulate_ppc() takes the
draws matrix and an optional design matrix to produce the
replicated data matrix.
y_rep <- simulate_ppc(
posterior_draws = posterior_draws,
X = X,
family = "gaussian",
sigma_posterior = sigma_draws
)
dim(y_rep) # 400 rows (draws) x 100 columns (observations)
#> [1] 400 100Each row of y_rep is one plausible dataset that the
model could have generated.
Running Posterior Predictive Diagnostics
ppc_diagnostics() computes several scalar discrepancy
statistics and returns an S3 object with a dedicated print
method.
y_obs <- example_data$y
diag_result <- ppc_diagnostics(y_obs = y_obs, y_rep = y_rep)
print(diag_result)
#>
#> -- Posterior Predictive Diagnostics (predictCheckR) --
#>
#> Draws : 400
#> Obs : 100
#>
#> Discrepancy Statistics:
#> Mean difference : -0.1263
#> Variance difference : -0.4878
#> Bayesian p-value : 0.2475
#> RMSE (pred vs. obs) : 1.025
#> Coverage (95% CI) : 0.94Interpreting the output:
| Statistic | Interpretation |
|---|---|
| Mean difference | Should be close to 0. |
| Variance difference | Should be close to 0. |
| Bayesian -value | Values near 0.5 indicate calibration. |
| RMSE | Lower is better; reflects point-prediction accuracy. |
| Coverage (95 % CI) | Should be near 0.95 for a well-calibrated model. |
Visualisation
Density Overlay
The density overlay is the workhorse visualisation for PPC. It superimposes the kernel density estimate of (dark line) over a random subsample of densities (light lines).
plot_ppc_overlay(
y_obs = y_obs,
y_rep = y_rep,
n_samples = 50
)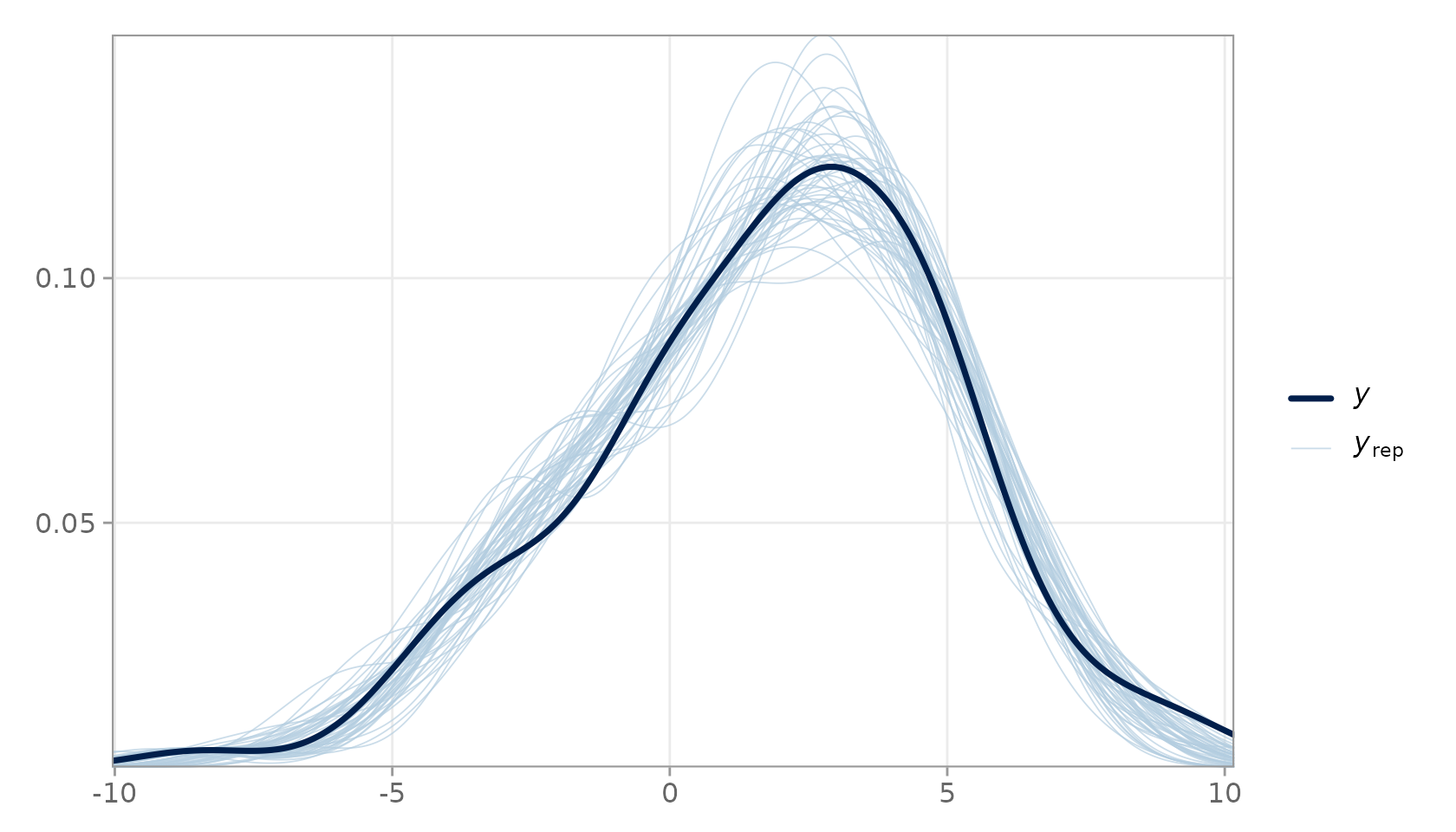
PPC density overlay. The dark line is the observed data; light lines are 50 random posterior predictive replicates.
Close agreement between the dark and light lines indicates that the model reproduces the marginal distribution of the response faithfully.
Test-Statistic Distribution
plot_ppc_stat() shows the distribution of
across all draws as a histogram, with a vertical line for
.
plot_ppc_stat(y_obs = y_obs, y_rep = y_rep, stat = "mean")
#> Note: in most cases the default test statistic 'mean' is too weak to detect anything of interest.
#> `stat_bin()` using `bins = 30`. Pick better value `binwidth`.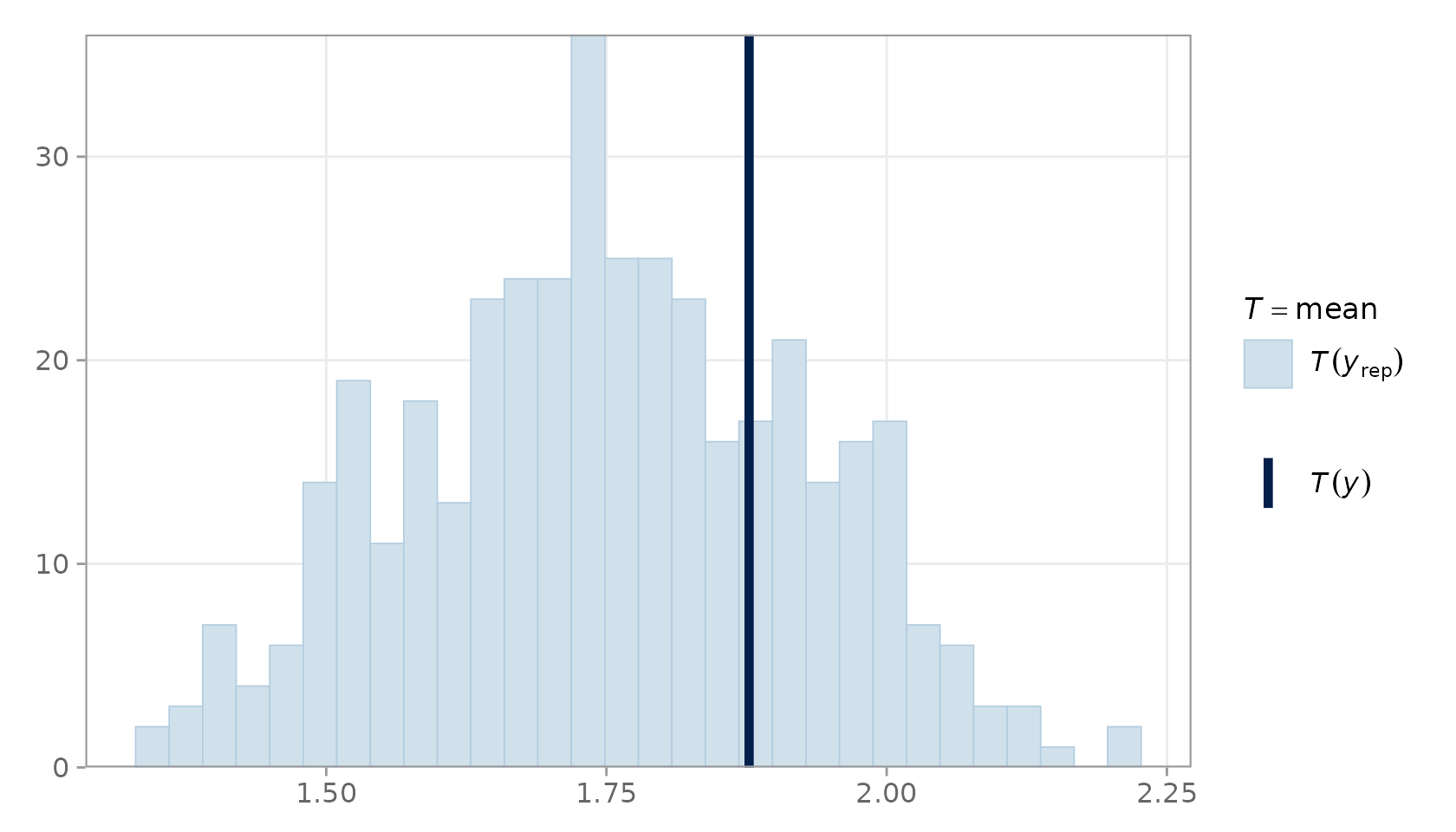
PPC statistic plot for the mean.
plot_ppc_stat(y_obs = y_obs, y_rep = y_rep, stat = "sd")
#> `stat_bin()` using `bins = 30`. Pick better value `binwidth`.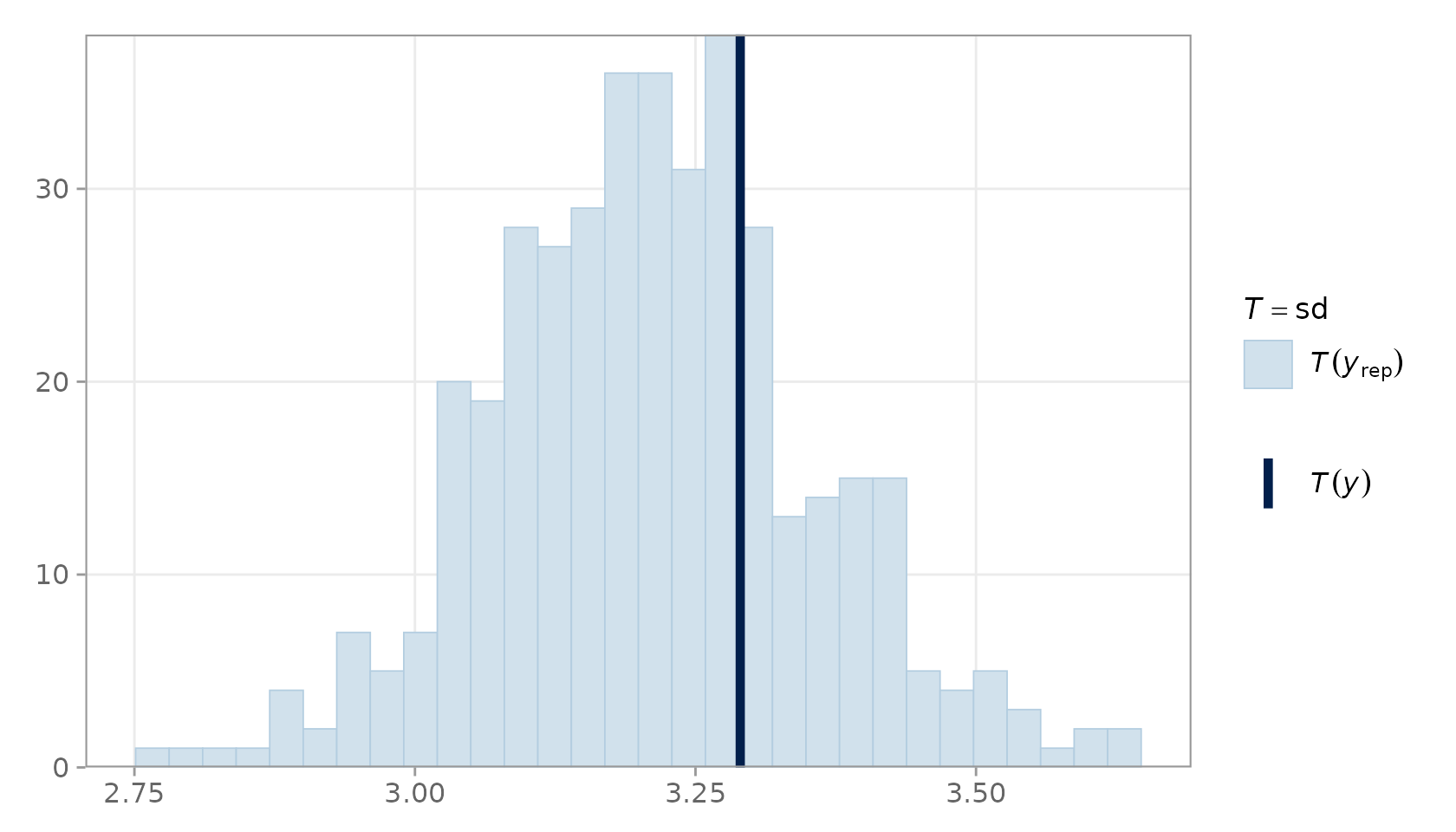
PPC statistic plot for the standard deviation.
The vertical line (observed statistic) should fall well within the histogram mass when the model is well-specified.
Model Comparison
compare_models_ppc() produces a side-by-side table of
predictive performance metrics for two competing models, making it
straightforward to assess which model better describes the data.
# Mis-specified model: draws centred on a shifted mean
set.seed(202)
posterior_draws_bad <- cbind(
intercept = rnorm(S, mean = 4.0, sd = 0.15), # wrong intercept
slope = rnorm(S, mean = 3.0, sd = 0.12)
)
y_rep_bad <- simulate_ppc(
posterior_draws = posterior_draws_bad,
X = X,
family = "gaussian",
sigma_posterior = sigma_draws
)
compare_models_ppc(
y_obs = y_obs,
y_rep1 = y_rep,
y_rep2 = y_rep_bad,
model_names = c("Correct Model", "Shifted Intercept")
)
#> metric Correct Model Shifted Intercept diff_m1_minus_m2
#> 1 RMSE 1.024788 2.120722 -1.095933
#> 2 MAE 0.828283 1.882697 -1.054414
#> 3 Pred. Variance Gap 9.792874 9.770498 0.022376The correctly-specified model should show lower RMSE and MAE, and a predictive variance gap close to zero.
Conclusion and Interpretation
Posterior predictive checking provides a transparent, simulation-based window into model adequacy. The predictCheckR workflow consists of four steps:
- Fit a Bayesian model and extract posterior draw vectors.
-
Simulate posterior predictive replicates with
simulate_ppc(). -
Diagnose discrepancies with
ppc_diagnostics(). -
Visualise with
plot_ppc_overlay()andplot_ppc_stat().
When the model is well-specified, replicated data should be statistically indistinguishable from the observed data. Systematic departures — heavy tails, multimodality, bias in a test statistic — point directly to the aspects of the data-generating process that the model fails to capture.
Session Information
sessionInfo()
#> R version 4.5.3 (2026-03-11)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.2 predictCheckR_0.1.0
#>
#> loaded via a namespace (and not attached):
#> [1] Matrix_1.7-4 bayesplot_1.15.0 gtable_0.3.6 jsonlite_2.0.0
#> [5] dplyr_1.2.1 compiler_4.5.3 Rcpp_1.1.1 tidyselect_1.2.1
#> [9] stringr_1.6.0 jquerylib_0.1.4 splines_4.5.3 systemfonts_1.3.2
#> [13] scales_1.4.0 textshaping_1.0.5 yaml_2.3.12 fastmap_1.2.0
#> [17] lattice_0.22-9 plyr_1.8.9 R6_2.6.1 labeling_0.4.3
#> [21] generics_0.1.4 knitr_1.51 tibble_3.3.1 desc_1.4.3
#> [25] bslib_0.10.0 pillar_1.11.1 RColorBrewer_1.1-3 rlang_1.1.7
#> [29] stringi_1.8.7 cachem_1.1.0 xfun_0.57 fs_2.0.1
#> [33] sass_0.4.10 S7_0.2.1 cli_3.6.5 pkgdown_2.2.0
#> [37] withr_3.0.2 magrittr_2.0.4 mgcv_1.9-4 digest_0.6.39
#> [41] grid_4.5.3 lifecycle_1.0.5 nlme_3.1-168 vctrs_0.7.2
#> [45] evaluate_1.0.5 glue_1.8.0 farver_2.1.2 ragg_1.5.2
#> [49] reshape2_1.4.5 rmarkdown_2.31 tools_4.5.3 pkgconfig_2.0.3
#> [53] htmltools_0.5.9