Bayesian Posterior Predictive Checking Utilities for R
Overview
predictCheckR provides a clean, minimal toolkit for Bayesian posterior predictive checking (PPC) built on the bayesplot and ggplot2 ecosystems.
Posterior predictive checking is the principled practice of simulating replicated data from the fitted model and comparing those simulations to the observed data. Systematic discrepancies reveal aspects of the data-generating process that the model fails to capture — before any information criterion is consulted.
The core workflow has four steps:
fit model → simulate_ppc() → ppc_diagnostics() → plot_ppc_overlay()Why Posterior Predictive Checking Matters
Standard goodness-of-fit metrics (AIC, BIC, WAIC) summarise the overall log-likelihood but do not tell you how a model fails. Predictive checking places simulated and observed data side by side, making model deficiencies immediately visible:
- Heavy tails not captured by a Gaussian likelihood
- Bimodality missed by a unimodal prior structure
- Systematic mean bias in a regression model
- Under- or overdispersion in count data
Installation
Install the development version directly from GitHub:
# install.packages("devtools")
devtools::install_github("utkarshpawade/predictCheckR", build_vignettes = TRUE)Quick Example
library(predictCheckR)
# Load the bundled example dataset (n = 100, y = 2 + 3x + N(0,1))
data(example_data)
# ── Step 1: Simulate fake posterior draws ──────────────────────────────────
set.seed(42)
S <- 400
n <- nrow(example_data)
posterior_draws <- cbind(
intercept = rnorm(S, mean = 2.0, sd = 0.15),
slope = rnorm(S, mean = 3.0, sd = 0.12)
)
X <- cbind(1, example_data$x)
sigma_draws <- abs(rnorm(S, mean = 1.0, sd = 0.08))
# ── Step 2: Generate posterior predictive samples ──────────────────────────
y_rep <- simulate_ppc(
posterior_draws = posterior_draws,
X = X,
family = "gaussian",
sigma_posterior = sigma_draws
)
# ── Step 3: Compute diagnostic statistics ──────────────────────────────────
diag <- ppc_diagnostics(y_obs = example_data$y, y_rep = y_rep)
print(diag)
#
# -- Posterior Predictive Diagnostics (predictCheckR) --
#
# Draws : 400
# Obs : 100
#
# Discrepancy Statistics:
# Mean difference : 0.0312
# Variance difference : -0.0241
# Bayesian p-value : 0.5425
# RMSE (pred vs. obs) : 0.1084
# Coverage (95% CI) : 0.96
# ── Step 4: Visualise ──────────────────────────────────────────────────────
plot_ppc_overlay(example_data$y, y_rep, n_samples = 50)
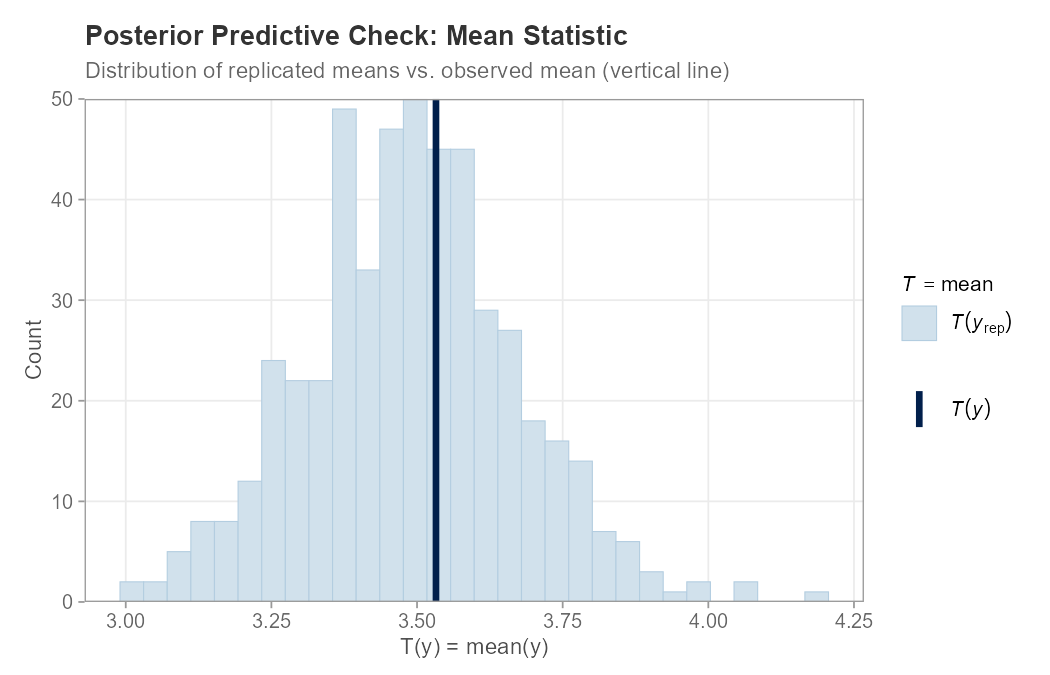
plot_ppc_stat(example_data$y, y_rep, stat = "mean")
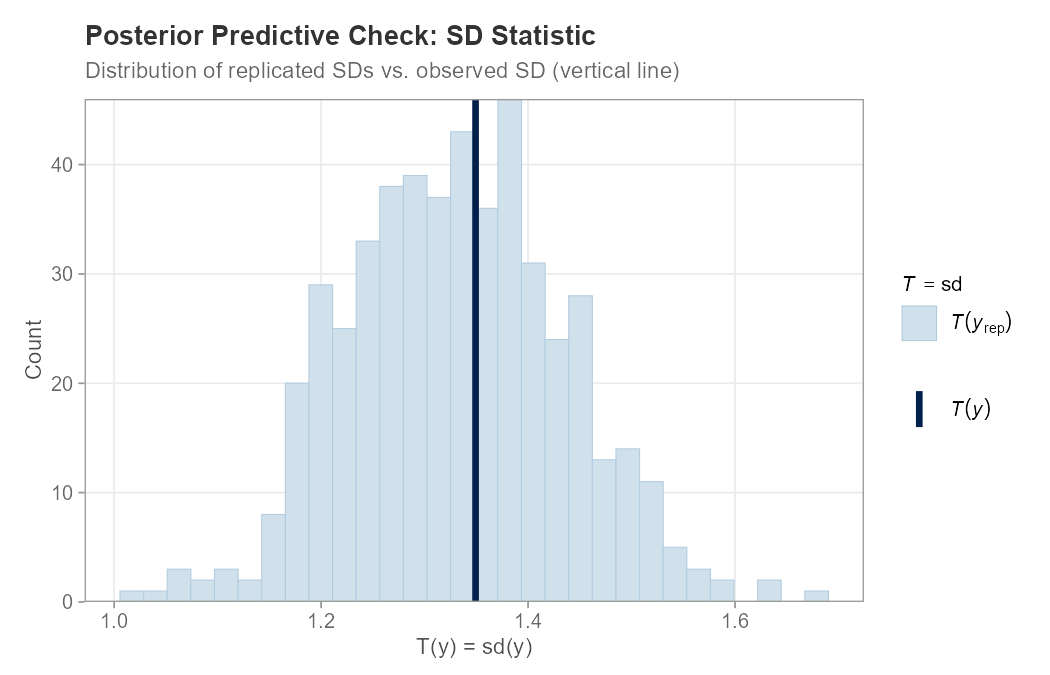
plot_ppc_stat(example_data$y, y_rep, stat = "sd")Example: Posterior Predictive Checking
The following example fits a simple Gaussian regression model, generates posterior predictive samples, runs diagnostics, and produces three plots. See example_ppc.R for the full reproducible script.
library(predictCheckR)
set.seed(42)
# Simulate data: y = 2 + 3x + N(0,1)
n <- 100
x <- seq(0, 1, length.out = n)
y_obs <- 2 + 3 * x + rnorm(n, mean = 0, sd = 1)
# Posterior draws from a well-specified model
S <- 500
posterior_draws <- cbind(intercept = rnorm(S, 2.0, 0.15),
slope = rnorm(S, 3.0, 0.12))
X <- cbind(1, x)
sigma_draws <- abs(rnorm(S, 1.0, 0.08))
# Generate posterior predictive samples
y_rep <- simulate_ppc(posterior_draws, X = X,
family = "gaussian", sigma_posterior = sigma_draws)
# Diagnostic statistics
ppc_diagnostics(y_obs, y_rep)
# Plots
plot_ppc_overlay(y_obs, y_rep, n_samples = 50)
plot_ppc_stat(y_obs, y_rep, stat = "mean")
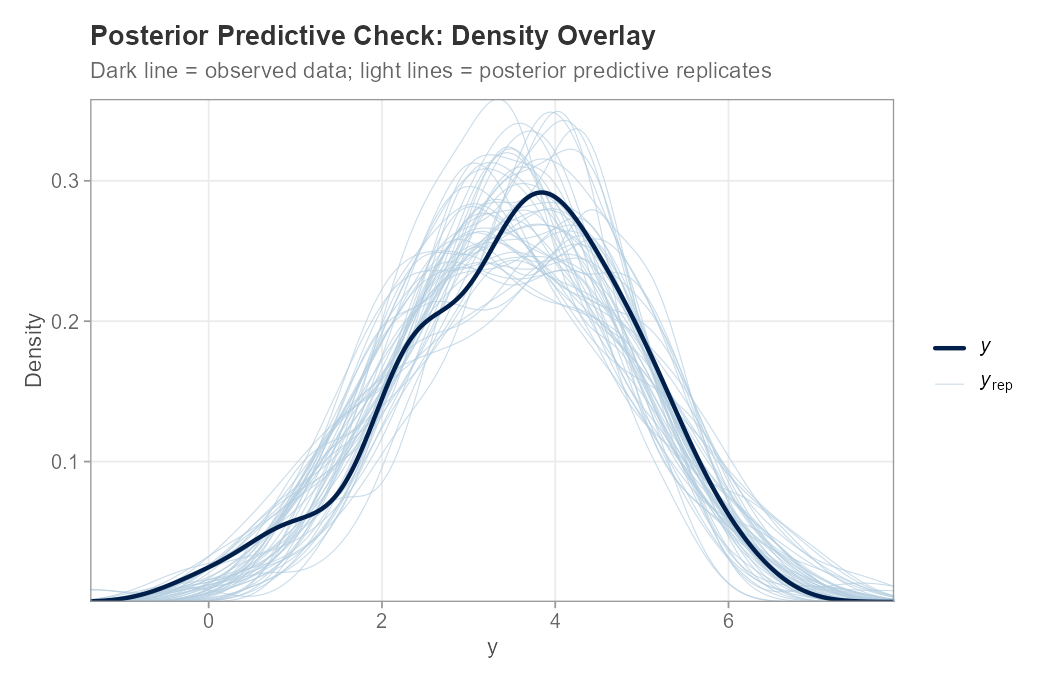
plot_ppc_stat(y_obs, y_rep, stat = "sd")PPC Density Overlay
The dark line shows the observed data distribution; the light lines are 50 randomly selected posterior predictive replicates. Good overlap indicates the model captures the marginal distribution of y well.
Function Reference
| Function | Purpose |
|---|---|
simulate_ppc() |
Generate posterior predictive sample matrix |
ppc_diagnostics() |
Compute mean diff, variance diff, Bayesian -value, RMSE, coverage |
print.ppc_diagnostics() |
Formatted S3 print method |
plot_ppc_overlay() |
Density overlay plot (wraps bayesplot::ppc_dens_overlay) |
plot_ppc_stat() |
Test-statistic distribution plot (wraps bayesplot::ppc_stat) |
compare_models_ppc() |
Side-by-side predictive performance table (RMSE, MAE, variance gap) |
theme_ppc() |
Clean ggplot2 theme for publication-quality PPC figures |
Model Comparison
# Competing model with mis-specified intercept
posterior_bad <- cbind(
intercept = rnorm(S, mean = 4.0, sd = 0.15),
slope = rnorm(S, mean = 3.0, sd = 0.12)
)
y_rep_bad <- simulate_ppc(posterior_bad, X = X,
sigma_posterior = sigma_draws)
compare_models_ppc(
y_obs = example_data$y,
y_rep1 = y_rep,
y_rep2 = y_rep_bad,
model_names = c("Correct", "Shifted")
)
# metric Correct Shifted diff_m1_minus_m2
# 1 RMSE 0.1084 1.9872 -1.8788
# 2 MAE 0.0871 1.9856 -1.8985
# 3 Pred. Variance Gap 0.0023 0.0018 0.0005Vignette
A detailed workflow vignette is included:
vignette("predictCheckR_workflow", package = "predictCheckR")Topics covered:
- Introduction to the posterior predictive distribution
- Mathematical derivation of Bayesian -values
- Full worked example using
example_data - Interpreting density overlays and test-statistic plots
- Model comparison workflow
Compatibility
predictCheckR works with any source of posterior draws represented as a numeric matrix:
-
brms — extract draws with
posterior::as_draws_matrix() -
rstan — extract draws with
rstan::extract(fit, permuted = FALSE) -
cmdstanr — use
fit$draws(format = "matrix") - Simulated draws for unit testing and tutorials
Citation
If you use predictCheckR in academic work, please cite:
predictCheckR Maintainer (2026). predictCheckR: Bayesian Posterior Predictive
Checking Utilities. R package version 0.1.0.
https://github.com/utkarshpawade/predictCheckRBibTeX:
@Manual{predictCheckR,
title = {predictCheckR: Bayesian Posterior Predictive Checking Utilities},
author = {{predictCheckR Maintainer}},
year = {2026},
note = {R package version 0.1.0},
url = {https://github.com/utkarshpawade/predictCheckR}
}License
MIT © 2026 predictCheckR Maintainer. See LICENSE.md.